Tenured Associate Professor
University of Florida J. Crayton Pruitt Family Department of Biomedical Engineering


We give priority to admit 1) top-ranking students from top universities, 2) world-level programming competition/challenge winners, and 3) students who published high-quality papers in this field.
Postdoc Position Open: A fully funded Post-doc positions are open on research in Medical Artificial Intelligence (MAI), brain-inspired AI, and AI for precision brain health.
Ph.D. Position Open: Multiple fully-funded Ph.D. positions are open on research in generative AI, foundation models, LLM, and digital twins.
Please check out the "Openings" tab to see more information.
An AI researcher in medicine and healthcare, Dr. Ruogu Fang is a tenured Associate Professor and Pruitt Family Endowed Faculty Fellow in the J. Crayton Pruitt Family Department of Biomedical Engineering at the University of Florida. Her research encompasses two principal themes: AI-empowered precision brain health and brain/bio-inspired AI. Her work involves addressing compelling questions, such as using machine learning techniques to quantify brain dynamics, facilitating early Alzheimer's disease diagnosis through novel imagery, predicting personalized treatment outcomes, designing precision interventions, and leveraging principles from neuroscience to develop the next-generation of AI. Dr. Fang is ranked among the Top 100 (top 1%) researchers worldwide in image analysis, according to ScholarGPS. Fang's current research is also rooted in the confluence of AI and multimodal medical image analysis. She is the PI of an NIH NIA RF1/R01, an NSF Research Initiation Initiative (CRII) Award, an NSF CISE IIS Award, a Ralph Lowe Junior Faculty Enhancement Award from Oak Ridge Associated Universities (ORAU). She has also received numerous recognitions. She was selected as the Rising Stars (Engineering) by the Academy of Science, Engineering, and Medicine of Florida (ASEMFL), the inaugural recipient of the Robin Sidhu Memorial Young Scientist Award from the Society of Brain Mapping and Therapeutics, an Best Paper Award from the IEEE International Conference on Image Processing, the University of Florida AI Course Award, an UF Herbert Wertheim College of Engineering Faculty Award for Excellence in Innovation, an UF BME Faculty Research Excellence Award and Faculty Teaching Excellence Award, among others. Fang's research has been featured by Forbes Magazine, The Washington Post, ABC, RSNA, and published in Lancet Digital Health, JAMA, PNAS, npj Digital Medicine, and npj Women Health. She also published in top conferences like ICCV, MICCAI, and She is an Associate Editor of the Journal Medical Image Analysis, a Topic Editor of Frontiers in Human Neuroscience, and a Guest Editor of CMIG. She is a reviewer for The Lancet, Nature Machine Intelligence, NatureScience Advances, etc. Her research has been supported by NSF, NIH, Oak Ridge Laboratory, DHS, DoD, NVIDIA, and the University of Florida. She is the President of Women in MICCAI (WiM) and an Associate Editor of the Medical Image Analysis journal. At the heart of her work is the Smart Medical Informatics Learning and Evaluation (SMILE) lab, where she is tirelessly dedicated to creating groundbreaking brain and neuroscience-inspired medical AI and deep learning models. The primary objective of these models is to comprehend, diagnose, and treat brain disorders, all while navigating the complexities of extensive and intricate datasets.






Brain dynamics, which reflects the healthy or pathological states of the brain with quantifiable, reproducible, and indicative dynamics values, remains the least understood and studied area of brain science despite its intrinsic and critical importance to the brain. Unlike other brain information such as the structural and sequential dimensions that have all been extensively studied with models and methods successfully developed, the 5th dimension, dynamics, has only very recently started receiving systematic analysis from the research community. The state-of-the-art models suffer from several fundamental limitations that have critically inhibited the accuracy and reliability of the dynamic parameters' computation. First, dynamic parameters are derived from each voxel of the brain spatially independently, and thus miss the fundamental spatial information since the brain is connected? Second, current models rely solely on single-patient data to estimate the dynamic parameters without exploiting the big medical data consisting of billions of patients with similar diseases.
This project aims to develop a framework for data-driven brain dynamics characterization, modeling and evaluation that includes the new concept of a 5th dimension – brain dynamics – to complement the structural 4-D brain for a complete picture. The project studies how dynamic computing of the brain as a distinct problem from the image reconstruction and de-noising of convention models, and analyzes the impact of different models for the dynamics analysis. A data-driven, scalable framework will be developed to depict the functionality and dynamics of the brain. This framework enables full utilization of 4-D brain spatio-temporal data and big medical data, resulting in accurate estimations of the dynamics of the brain that are not reflected in the voxel-independent models and the single patient models. The model and framework will be evaluated on both simulated and real dual-dose computed tomography perfusion image data and then compared with the state-of-the-art methods for brain dynamics computation by leveraging collaborations with Florida International University Herbert Wertheim College of Medicine, NewYork-Presbyterian Hospital / Weill Cornell Medical College (WCMC) and Northwell School of Medicine at Hofstra University. The proposed research will significantly advance the state-of-the-art in quantifying and analyzing brain structure and dynamics, and the interplay between the two for brain disease diagnosis, including both the acute and chronic diseases. This unified approach brings together fields of Computer Science, Bioengineering, Cognitive Neuroscience and Neuroradiology to create a framework for precisely measuring and analyzing the 5th dimension – brain dynamics – integrated with the 4-D brain with three dimensions from spatial data and one dimension from temporal data. Results from the project will be incorporated into graduate-level multi-disciplinary courses in machine learning, computational neuroscience and medical image analysis. This project will open up several new research directions in the domain of brain analysis, and will educate and nurture young researchers, advance the involvement of underrepresented minorities in computer science research, and equip them with new insights, models and tools for developing future research in brain dynamics in a minority serving university.
Ph.D. in Electrical and Computer Engineering
Cornell University, Ithaca, NY.
B.E. in Information Engineering
Zhejiang University, Hangzhou, China
VSI: Foundation Models for Computational Pathology in Medical Image Analysis, The Engineering in Medicine and Biology Conference
Current Opinion in Biomedical Engineering, Artificial Intelligence in Biomedical Engineering
Women in MICCAI (WiM)
Medical Image Computing and Computer Assisted Intervention (MICCAI) Society
Women in Brain Imaging and Stimulation, Frontiers in Human Neuroscience Journal
MICCAI Conference
The International Conference on Medical Image Computing and Computer Assited Intervention (MICCAI)
Medical Image Analysis
Computerized Medical Imaging and Graphics, Elsevier
Nature Neuroscience, npj Digital Medicine, Scientific Data, IEEE Reviews in Biomedical Engineering, The Lancet, Nature Machine Intelligence, Science Advances, IEEE TPAMI, IEEE TNNLS, MIA, IEEE TMI, IEEE TIP
University of Florida J. Crayton Pruitt Family Department of Biomedical Engineering
Stanford University School of Medicine
Stanford University School of Humanities and Sciences
Harvard University
Harvard Medical School
Massachusetts General Brigham
University of Florida UF Intelligent Critical Care Center (IC3)
University of Florida Department of Electrical and Computer Engineering
University of Florida Department of Radiology, College of Medicine
University of Florida Department of Computer & Information Science & Engineering
University of Florida The Center for Cognitive Aging and Memory Clinical Translational Research (CAM)
University of Florida Genetics Institute (UFGI)
University of Florida UF Health Cancer Center
University of Florida Emerging Pathogens Institute
University of Florida Norman Fixel Institute for Neurological Diseases

















































































































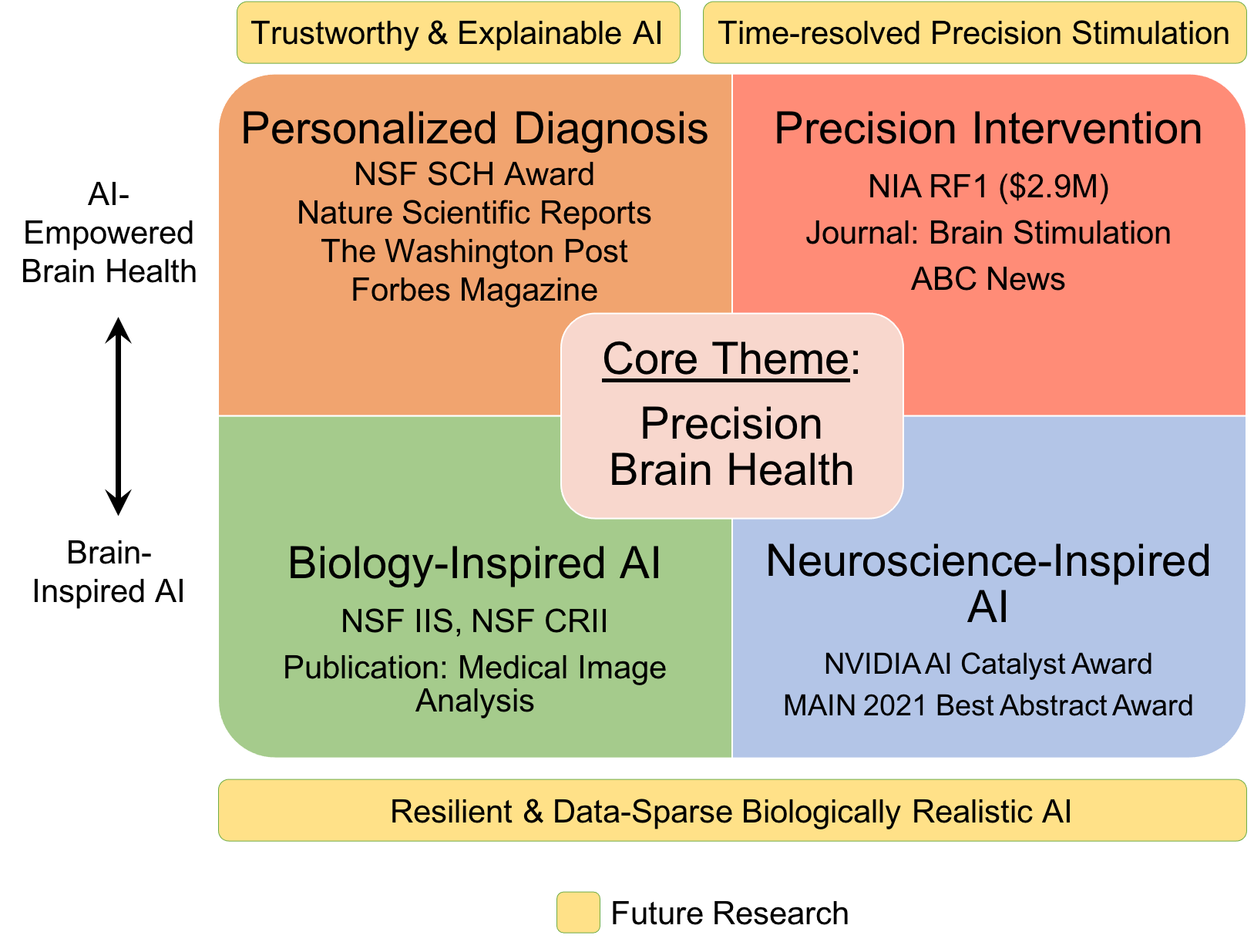
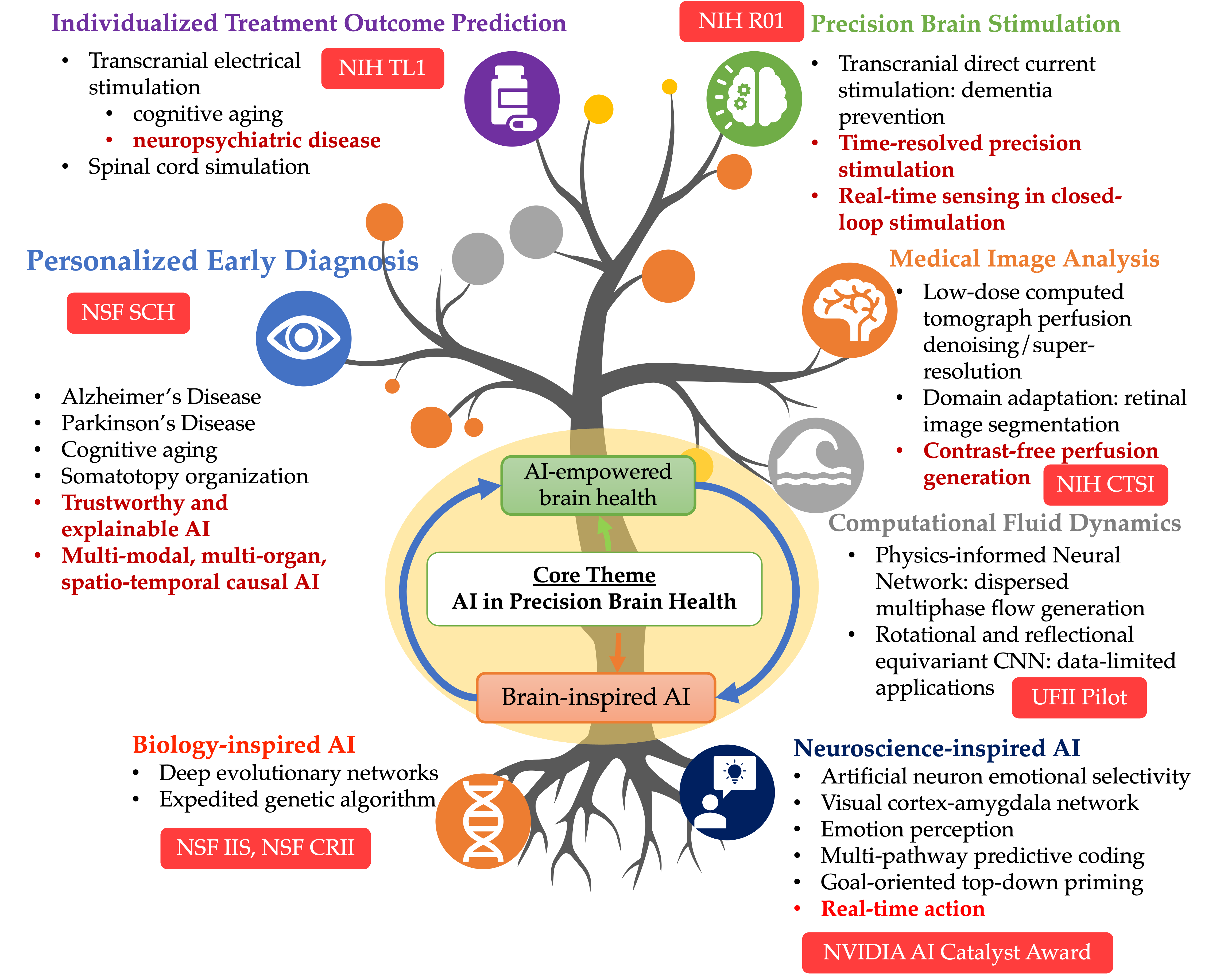

The University of Florida (UF) proposes to develop a nation-wide digital course to upskill biomedical researchers’ expertise in artificial intelligence (AI). The AI Passport for Biomedical Research (AIPassportBMR) overall objectives are to provide diverse biomedical researchers the opportunity to augment their skills in a concise and comprehensive course based on innovative learning techniques and will be the first scalable, self-correcting AI training program that is dynamic enough to withstand constant evolution of AI technologies. AIPassportBMR will focus on technical skills while providing mentorship on how to expand biomedical research into the AI field with highly collaborative sessions that include national leaders in medical AI. Moreover, due to the disparity in the current AI workforce, participants will be selected to increase diversity and inclusive excellence. Biomedical research and AI each align with institutional priorities, possess robust resources, and represent highly collaborative and successful scientific communities at UF. Designed as a digital experiential learning community, this program will enable predoctoral trainees and early-stage investigators in biomedical, behavioral, and clinical sciences to acquire the multidisciplinary skills necessary to integrate AI into their research while creating a nationwide community of likeminded mentors and peers. Our approach aligns with the objectives outlined in the IPERT initiate to provide skills development and mentoring to an inclusive audience through the following three overarching aims: Aim 1. AIPassportBMR Program: Develop the educational and technological infrastructure for community-driven experiential biomedical AI training using a See-Practice-Share-Reflect learning approach, Aim 2. Real-World Evaluation: Implement, evaluate, and fine-tune AIPassportBMR using an implementation mapping framework and Cognitive Theory of Culture to align instructional design with learners’ subcultural and educational needs, Aim 3. AI Digital Community of Practice: Build a support mosaic network of peers, mentors and coaches and the capacity for sustainable nationwide dissemination of the AI research training program. To further expand AIPassportBMR, the digital community learning platform program will be disseminated nationwide to build capacity for biomedical AI workforce development and support replication for all biomedical researchers. This program is committed to promoting diversity and inclusive excellence for its participants.


Human emotions are dynamic, multidimensional responses to challenges and opportunities that emerge from network interactions in the brain. Disruptions of these dynamics underlie emotional dysregulation in many mental disorders including anxiety and depression. To empirically study the neural basis of human emotion inference, experimenters often have observers view natural images varying in affective content, while at the same time recording their brain activity using electroencephalogram (EEG) and/or Functional magnetic resonance imaging (fMRI). Despite extensive research over the last few decades, much remains to be learned about the computational principles subserving the recognition of emotions in natural scenes. A major roadblock faced by empirical neuroscientists is the inability to carry out precisely manipulate human neural systems and test the consequences in imaging data. Deep Neural Networks (DNN), owing to their high relevance to human neural systems and extraordinary prediction capability, have become a promising tool for testing these sorts of hypotheses in swift and nearly costless computer simulations. The overarching goal of this project is to develop a neuroscience-inspired, DNN-based deep learning framework for emotion inference in real-world scenarios by synergistically integrating neuron-, circuit-, and system-level mechanisms. Recognizing that the state-of-the-art DNNs are centered on bottom-up and feedforward-only processing, which disagrees with the strong goal-oriented top-down modulation recurrence observed in the physiology, this project aims to enrich DNNs and enable closer AI-neuroscience interaction by incorporating goal-oriented top-down modulation and reciprocal interactions DNNs and test the model assumptions and predictions on neuroimaging data. To meet these goals, the project aims to develop a brain-inspired goal-oriented and bidirectional deep learning model for emotion inference. Despite the great promise shown by today?s deep learning as a framework for modeling biological vision, their architecture is limited to emulating the visual cortex for face/object/scene recognition and rarely goes beyond the inferotemporal cortex (IT), which is necessary for modeling high-level cognitive processes. In this project, we propose to build a biologically plausible deep learning architecture by integrating an in-silico amygdala module into the visual cortex architecture in DNN (the VCA model). The researchers hope to build neuron-, circuit-, and system-level modulation via goal-oriented attention priming, and multi-pathway predictive coding to 1) elucidate the mechanism of selectivity underlying preference and response to naturalistic emotions by artificial neurons; 2) differentiate fine-grained emotional responses via multi-path predictive coding, and 3) refine the neuroscientific understanding of human neuro-behavioral data by comparing attention priming and temporal generalization observed in simultaneous fMRI-EEG data to the computational observations using our brain-inspired VCA model. This project introduces two key innovations, both patterned after how brain operates, into DNN architecture and demonstrate their superior performance when applied to complex real-world tasks. Successful execution of the project can lead to the development of a new generation of AI-models that are inspired by neuroscience and that may in turn power neuroscience research.

There is a pressing need for effective interventions to remediate age-related cognitive decline and alter the trajectory toward Alzheimer’s disease. The NIA Alzheimer’s Disease Initiative funded Phase III Augmenting Cognitive Training in Older Adults (ACT) trial aimed to demonstrate that transcranial direct current stimulation (tDCS) paired with cognitive training could achieve this goal. The present study proposes a state of the art secondary data analysis of ACT trial data that will further this aim by 1) elucidate mechanism of action underlying response to tDCS treatment with CT, 2) address heterogeneity of response in tDCS augmented CT by determining how individual variation in the dose of electrical current delivered to the brain interacts with individual brain anatomical characteristics; and 3) refine the intervention strategy of tDCS paired with CT by evaluating methods for precision delivery targeted dosing characteristics to facilitate tDCS augmented outcomes. tDCS intervention to date, including ACT, apply a fixed dosing approach whereby a single stimulation intensity (e.g., 2mA) and set of electrode positions on the scalp (e.g., F3/F4) is applied to all participants/patients. However, our recent work has demonstrated that age-related changes in neuroanatomy as well as individual variability in head/brain structures (e.g., skull thickness) significantly impacts the distribution and intensity of electrical current induced in the brain from tDCS. This project will use person-specific MRI-derived finite element computational models of electric current characteristics (current intensity and direction of current flow) and new methods for enhancing the precision and accuracy of derived models to precisely quantify the heterogeneity of current delivery in older adults. We will leverage these individualized precision models with state-of-the-art support vector machine learning methods to determine the relationship between current characteristics and treatment response to tDCS and CT. We will leverage the inherent heterogeneity of neuroanatomy and fixed current delivery to provide insight in the not only which dosing parameters were associated with treatment response, but also brain region specific information to facilitate targeted delivery of stimulation in future trials. Further still, the current study will also pioneer new methods for calculation of precision dosing parameters for tDCS delivery to potentially optimize treatment response, as well as identify clinical and demographic characteristics that are associated with response to tDCS and CT in older adults. Leveraging a robust and comprehensive behavioral and multimodal neuroimaging data set for ACT with advanced computational methods, the proposed study will provide critical information for mechanism, heterogeneity of treatment response and a pathway to refined precision dosing approaches for remediating age- related cognitive decline and altering the trajectory of older adults toward Alzheimer’s disease.

There is a pressing need for effective interventions to remediate age-related cognitive decline and alter the trajectory toward Alzheimer’s disease. The NIA Alzheimer’s Disease Initiative funded Phase III Augmenting Cognitive Training in Older Adults (ACT) trial aimed to demonstrate that transcranial direct current stimulation (tDCS) paired with cognitive training could achieve this goal. The present study proposes a state of the art secondary data analysis of ACT trial data that will further this aim by 1) elucidate mechanism of action underlying response to tDCS treatment with CT, 2) address heterogeneity of response in tDCS augmented CT by determining how individual variation in the dose of electrical current delivered to the brain interacts with individual brain anatomical characteristics; and 3) refine the intervention strategy of tDCS paired with CT by evaluating methods for precision delivery targeted dosing characteristics to facilitate tDCS augmented outcomes. tDCS intervention to date, including ACT, apply a fixed dosing approach whereby a single stimulation intensity (e.g., 2mA) and set of electrode positions on the scalp (e.g., F3/F4) is applied to all participants/patients. However, our recent work has demonstrated that age-related changes in neuroanatomy as well as individual variability in head/brain structures (e.g., skull thickness) significantly impacts the distribution and intensity of electrical current induced in the brain from tDCS. This project will use person-specific MRI-derived finite element computational models of electric current characteristics (current intensity and direction of current flow) and new methods for enhancing the precision and accuracy of derived models to precisely quantify the heterogeneity of current delivery in older adults. We will leverage these individualized precision models with state-of-the-art support vector machine learning methods to determine the relationship between current characteristics and treatment response to tDCS and CT. We will leverage the inherent heterogeneity of neuroanatomy and fixed current delivery to provide insight in the not only which dosing parameters were associated with treatment response, but also brain region specific information to facilitate targeted delivery of stimulation in future trials. Further still, the current study will also pioneer new methods for calculation of precision dosing parameters for tDCS delivery to potentially optimize treatment response, as well as identify clinical and demographic characteristics that are associated with response to tDCS and CT in older adults. Leveraging a robust and comprehensive behavioral and multimodal neuroimaging data set for ACT with advanced computational methods, the proposed study will provide critical information for mechanism, heterogeneity of treatment response and a pathway to refined precision dosing approaches for remediating age- related cognitive decline and altering the trajectory of older adults toward Alzheimer’s disease.

Driven by its performance accuracy, machine learning (ML) has been used extensively for various applications in the healthcare domain. Despite its promising performance, researchers and the public have grown alarmed by two unsettling deficiencies of these otherwise useful and powerful models. First, there is a lack of trustworthiness - ML models are prone to interference or deception and exhibit erratic behaviors when in action dealing with unseen data, despite good practice during the training phase. Second, there is a lack of interpretability - ML models have been described as 'black-boxes' because there is little explanation for why the models make the predictions they do. This has called into question the applicability of ML to decision-making in critical scenarios such as image-based disease diagnostics or medical treatment recommendation. The ultimate goal of this project is to develop computational foundation for trustworthy and explainable Artificial Intelligence (AI), and offer a low-cost and non-invasive ML-based approach to early diagnosis of neurodegenerative diseases. In particular, the project aims to develop computational theories, ML algorithms, and prototype systems. The project includes developing principled solutions to trustworthy ML and making the ML prediction process transparent to end-users. The later will focus on explaining how and why an ML model makes such a prediction, while dissecting its underlying structure for deeper understanding. The proposed models are further extended to a multi-modal and spatial-temporal framework, an important aspect of applying ML models to healthcare. A verification framework with end-users is defined, which will further enhance the trustworthiness of the prototype systems. This project will benefit a variety of high-impact AI-based applications in terms of their explainability, trustworthy, and verifiability. It not only advances the research fronts of deep learning and AI, but also supports transformations in diagnosing neurodegenerative diseases.
This project will develop the computational foundation for trustworthy and explainable AI with several innovations. First, the project will systematically study the trustworthiness of ML systems. This will be measured by novel metrics such as, adversarial robustness and semantic saliency, and will be carried out to establish the theoretical basis and practical limits of trustworthiness of ML algorithms. Second, the project provides a paradigm shift for explainable AI, explaining how and why a ML model makes its prediction, moving away from ad-hoc explanations (i.e. what features are important to the prediction). A proof-based approach, which probes all the hidden layers of a given model to identify critical layers and neurons involved in a prediction from a local point of view, will be devised. Third, a verification framework, where users can verify the model's performance and explanations with proofs, will be designed to further enhance the trustworthiness of the system. Finally, the project also advances the frontier of neurodegenerative diseases early diagnosis from multimodal imaging and longitudinal data by: (i) identifying retinal vasculature biomarkers using proof-based probing in biomarker graph networks; (ii) connecting biomarkers of the retina and the brain vasculature via cross- modality explainable AI model; and, (iii) recognizing the longitudinal trajectory of vasculature biomarkers via a spatio-temporal recurrent explainable model. This synergistic effort between computer science and medicine will enable a wide range of applications to trustworthy and explainable AI for healthcare. The results of this project will be assimilated into the courses and summer programs that the research team have developed with specially designed projects to train students with trustworthy and explainable AI.

History has taught us that exposures to radionuclides can happen any day almost anywhere in the US and elsewhere and we have done little to prepare ourselves. Our ability to perform dosimetry modeling for such scenarios and efforts into biomarker and mitigation discovery are archaic and our tendency to rely on external beam radiation to model these is utterly misplaced. We should and we can do much better. This program centers on the hypothesis that radiation from internal emitters is very unevenly distributed within a body, amongst organs, and even within organs, tissues and cells. The half-life and decay schema of the radionuclide, its activity and concentration, particle size and morphology, and its chemical form and solubility are all critical, as are the route of uptake, tissue structure, genetic makeup, physiology, danger signaling and the crosstalk with the immune system. Conceptually this suggests that the analysis of radionuclide distribution requires measurements at the MESO, MICRO and NANO level for accurate dosimetry modeling and biokinetics analyses, that will much better align with biological endpoints, and therefore with meaningful countermeasure development. In many ways our program integrates the three main pillars of radiation science, namely radiation physics, radiation chemistry and radiation biology, taking into account pharmacokinetics and pharmacodynamics aspects of particle distribution at subcellular, cellular, and tissue levels. In other words, to understand the biological effects of internal emitters and find the best possible mitigation strategies a systematic study is called for, one that includes but is not limited to: a) radionuclide physical and chemical form and intravital migration, b) protracted exposure times, c) radiation quality parameters, d) novel virtual phantom modeling beyond few MACRO reference models ; e) novel biokinetics with sex- and age- specificity; f) MESO, MICRO and NANO scale histology and immunohistochemistry with integrated radionuclide distribution information; g) exploration of molecular biomarkers of radionuclide intake and contamination and h) countermeasures that modulate radionuclide distribution and possibly also improve DNA, cell and tissue repair. We have assembled a team with diverse scientific expertise that can tackle these challenges within an integrated program. There is an incredibly impressive technological toolbox at our disposal and our goal is to generate a meaningful blueprint for understanding and predicting biological consequences of exposure to radionuclides. The possible benefits of this program to the radiation research community and the general population are immense.

The purpose of the SHARE P01 research program project is to address HIV and alcohol use around three themes; 1) Emerging adulthood (ages 18 -29); 2) Self-management of HIV and alcohol; and 3) Translational behavioral science. Emerging adulthood is a developmental stage marked by significant change in social roles, expectations as a new adult, and increased responsibilities. It is also marked by poor HIV self- management and increased alcohol use. Emerging adults with HIV (hereafter called young people living with HIV; YPLWH) may face even more challenges given intersectional stigma. This age group continues to have very high rates of new HIV infections. Interventions designed specifically for the unique developmental challenges of emerging adults are needed, yet emerging adults are often included with older adults in intervention programs. The concept of self-management emerged concurrently within both the substance abuse and chronic illness literatures, and fits well with the developmental challenges of emerging adulthood. Self-management, a framework we have utilized in our work with YPLWH, refers to the ability to manage symptoms, treatments, lifestyle changes, and consequences of health conditions. Current research now identifies individual-level self-management skills such as self-control, decision-making, self-reinforcement, and problem solving as that protect against substance use and improve other health outcomes and can be embedded in the Information-Motivation-Behavioral Skills model. Although we have conducted multiple studies with YPLWH, only one intervention to date (Healthy Choices conducted by our team) improved both alcohol use and viral suppression in YPLWH in large trials. The goal of the SHARE P01 is to utilize advances in translational behavioral science to optimize behavioral interventions and define new developmentally- and culturally-appropriate intervention targets to improve self-management of alcohol and HIV in YPLWH. We will focus our efforts in Florida, a state hardest hit by the HIV epidemic but with a particularly strong academic- community partnership to support translation. We have assembled research teams to conduct self- management studies across the translational spectrum to address self-management and improve alcohol use and viral suppression (and thereby reduce transmission) in diverse YPLWH in Florida. The P01 will consist of three research projects (DEFINE, ENGAGE, and SUSTAIN), representing different stages on the translational spectrum and targeting different core competencies, supported by two cores (Community Engagement Core and Data Science Core). If successful, the SHARE P01 has the potential to greatly advance programs promoting self-management of HIV and alcohol use among a particularly vulnerable, but under-researched group, emerging adults living with HIV. SHARE also has a high potential for scale-up and implementation beyond Florida and across the United States.

As persons living with HIV (PLWH) live longer, approximately 50% will experience HIV-related cognitive dysfunction, which may affect daily activities, contribute to morbidity and mortality, and increase the likelihood of HIV transmission. Alcohol consumption among PLWH may further exacerbate long-term cognitive dysfunction, with the presumed mechanism involving the gut microbiome, microbial translocation, systemic inflammation, and ultimately neuroinflammation. However, there are many gaps in our understanding regarding the specific pathophysiological mechanisms, and a need to offer interventions that are effective and acceptable in helping PLWH to reduce drinking or to protect them against alcohol-related harm. The overarching goal of this P01 is to identify and ultimately implement new/improved, targeted interventions that will improve outcomes related to cognitive and brain dysfunction in persons with HIV who drink alcohol. The proposed P01 activity will extend our current line of research that forms the core of the Southern HIV & Alcohol Research Consortium (SHARC). The specific aims of this P01 are to: 1) improve our understanding of the specific mechanisms that connect the gut microbiome to cognitive and brain health outcomes in persons with HIV; 2) evaluate interventions that are intended to reduce the impact of alcohol on brain and cognitive health in persons with HIV; and 3) connect and extend the research activity from this P01 with the training programs and community engagement activity in the SHARC. Our P01 will utilize two cores that provide infrastructure to two Research Components (RC1, RC2). The two RC will together enroll 200 PLWH with at-risk drinking into clinical trials that share common timepoints and outcome assessments. RC1 will compare two strategies to extend contingency management to 60 days, using breathalyzers and wrist-worn biosensors to monitor drinking. RC2 uses a hybrid trial design to evaluate two biomedical interventions targeting the gut-brain axis. One intervention is a wearable, transcutaneous vagus nerve stimulator that is hypothesized to stimulate the autonomic nervous system, resulting in decreased inflammation and improved cognition. The other intervention is a probiotic supplement intended to improve the gut microbiome in persons with HIV and alcohol consumption. All participants in RC2, and a subset of those in RC1 will have neuroimaging at two timepoints. The Data Science Core will provide data management and analytical support, and will analyze existing data and the data collected from this P01 using a machine learning and AI approach to identify factors associated with intervention success or failure. The Administrative Core will provide scientific leadership, clinical research and recruitment infrastructure, and connection to the outstanding training programs, development opportunities, and community engagement provided by the SHARC. Our community engagement with diverse populations, and collection of acceptability data from clinical trial participants, will facilitate our readiness to scale up the most promising interventions and move towards implementation in the next phase of our research.

The number of persons living with HIV (PLWH) continues to increase in the United States. Alcohol consumption is a significant barrier to both achieving the goal of ending the HIV epidemic and preventing comorbidities among PLWH, as it contributes to both HIV transmission and HIV-related complications. Recent advances in data capture systems such as mHealth devices, medical imaging, and high-throughput biotechnologies make large/complex research and clinical datasets available, including survey data, multi-omics data, electronic medical records, and/or other sources of reliable information related to engagement in care. This offers tremendous potential of applying “big” data to extract knowledge and insights regarding fundamental physiology, understand the mechanisms by which the pathogenic effects of biotic and abiotic factors are realized, and identify potential intervention targets. We propose to integrate the disparate data sources maintained by our partners and then utilize the big data to address research questions in treating HIV and alcohol-related morbidity and mortality. Specifically, we will pursue the following three aims: 1) Integrate the disparate data sources through standardization, harmonization, and merging; 2) Develop a web-based data sharing platform including virtual data sharing communities, data privacy protection, streamlined data approval and access, and tracking of ongoing research activities; 3) Provide statistical support to junior investigators to use the data repository for exploratory data analysis and proposal development. The proposed study will tap into disparate data sources, unleash the potential of data and information, accelerate knowledge discovery, advance data-powered health, and transform discovery to improve health outcomes for PLWH.

Classical aversive conditioning is a well-established laboratory model for studying acquisition and extinction of defensive responses. In experimental animals, as well as in humans, research to date has been mainly focused on the role of limbic structures (e.g., the amygdala) in these responses. Recent evidence has begun to stress the important contribution by the brain’s sensory and attention control systems in maintaining the neural representations of conditioned responses and in facilitating their extinction. The proposed research breaks new ground by combining novel neuroimaging techniques with advanced computational methods to examine the brain’s visual and attention processes underlying fear acquisition and extinction in humans. Major advances will be made along three specific aims. In Aim 1, we characterize the brain network dynamics of visuocortical threat bias formation, extinction, and recall in a two-day learning paradigm. In Aim 2, we establish and test a computational model of threat bias generalization. In Aim 3, we examine the relation between individual differences in generalization and recall of conditioned visuocortical threat biases and individual differences in heightened autonomic reactivity to conditioned threat, a potential biomarker for assessing the predisposition to developing the disorders of fear and anxiety. It is expected that accomplishing these research aims will address two NIMH strategic priorities: defining the circuitry and brain networks underlying complex behaviors (Objective 1) and identifying and validating new targets for treatment that are derived from the understanding of disease mechanisms (Objective 3). It is further expected that this project will enable a paradigm shift in research on dysfunctional attention to threat from one that focuses primarily on limbic-prefrontal circuits to one that emphasizes the interactions among sensory, attention, executive control and limbic systems.

Across the globe, there has been a considerable growth in the number of people diagnosed with Parkinsonism. Estimates indicate that from 1990 to 2015 the number of Parkinsonism diagnoses doubled, with more than 6 million people currently carrying the diagnosis, and by year 2040, 12 and 14.2 million people will be diagnosed with Parkinsonism. Parkinson’s disease (PD), multiple system atrophy Parkinsonian variant (MSAp), and progressive supranuclear palsy (PSP) are neurodegenerative forms of Parkinsonism, which can be difficult to diagnose as they share similar motor and non-motor features, and they each have an increased chance of developing dementia. In the first five years of a PD diagnosis, about 58% of PD are misdiagnosed, and of these misdiagnoses about half have either MSA or PSP. Since PD, MSAp, and PSP require unique treatment plans and different medications, and clinical trials testing new medications require the correct diagnosis, there is an urgent need for both clinic ready and clinical-trial ready markers for differential diagnosis of PD, MSAp, and PSP. Over the past decade, we have developed diffusion imaging as an innovative biomarker for differentiating PD, MSAp, and PSP. In this proposal, we will leverage our extensive experience to create a web-based software tool that can process diffusion imaging data from anywhere in the world. We will disseminate and test the tool in the largest prospective cohort of participants with Parkinsonism (PD, MSAp, PSP), working closely with the Parkinson Study Group. The reason to test this in the Parkinson Study Group network, is because they are the community that evaluates Phase II and Phase III clinical trials in Parkinsonism. This web-based software tool will be capable of reading raw diffusion imaging data, performing quality assurance procedures, analyzing the data using a validated pipeline, and providing imaging metrics and diagnostic probability. We will test the performance of the wAID-P by enrolling 315 total subjects (105 PD, 105 MSAp, 105 PSP) across 21 sites in the Parkinson Study Group. Each site will perform imaging, clinical scales, diagnosis, and will upload the data to the web-based software tool. The clinical diagnosis will be blinded to the diagnostic algorithm and the imaging diagnosis will be compared to the movement disorders trained neurologist diagnosis. We will also enroll a portion of the cohort into a brain bank to ascertain pathological confirmation and to test the algorithm against cases with post-mortem diagnoses. The final outcome will be to disseminate a validated diagnostic algorithm to the Parkinson neurological and radiological community and to make it available to all on a website.

The temporal dynamics of blood flows through the network of cerebral arteries and veins provides a window into the health of the human brain. Since the brain is vulnerable to disrupted blood supply, brain dynamics serves as a crucial indicator for many kinds of neurological diseases such as stroke, brain cancer, and Alzheimer's disease. Existing efforts at characterizing brain dynamics have predominantly centered on 'isolated' models in which data from single-voxel, single-modality, and single-subject are characterized. However, the brain is a vast network, naturally connected on structural and functional levels, and multimodal imaging provides complementary information on this natural connectivity. Thus, the current isolated models are deemed not capable of offering the platform necessary to enable many of the potential advancements in understanding, diagnosing, and treating neurological and cognitive diseases, leaving a critical gap between the current computational modeling capabilities and the needs in brain dynamics analysis. This project aims to bridge this gap by exploiting multi-scale structural (voxel, vasculature, tissue) connectivity and multi-modal (anatomical, angiography, perfusion) connectivity to develop an integrated connective computational paradigm for characterizing and understanding brain dynamics.

The temporal dynamics of blood flows through the network of cerebral arteries and veins provides a window into the health of the human brain. Since the brain is vulnerable to disrupted blood supply, brain dynamics serves as a crucial indicator for many kinds of neurological diseases such as stroke, brain cancer, and Alzheimer's disease. Existing efforts at characterizing brain dynamics have predominantly centered on 'isolated' models in which data from single-voxel, single-modality, and single-subject are characterized. However, the brain is a vast network, naturally connected on structural and functional levels, and multimodal imaging provides complementary information on this natural connectivity. Thus, the current isolated models are deemed not capable of offering the platform necessary to enable many of the potential advancements in understanding, diagnosing, and treating neurological and cognitive diseases, leaving a critical gap between the current computational modeling capabilities and the needs in brain dynamics analysis. This project aims to bridge this gap by exploiting multi-scale structural (voxel, vasculature, tissue) connectivity and multi-modal (anatomical, angiography, perfusion) connectivity to develop an integrated connective computational paradigm for characterizing and understanding brain dynamics.

This project establishes the NSF Industry/University Collaborative Research Center for Big Learning (CBL). The vision is to create intelligence towards intelligence-driven society. Through catalyzing the fusion of diverse expertise from the consortium of faculty members, students, industry partners, and federal agencies, CBL seeks to create state-of-the-art deep learning methodologies and technologies and enable intelligent applications, transforming broad domains, such as business, healthcare, Internet-of-Things, and cybersecurity. This timely initiative creates a unique platform for empowering our next-generation talents with cutting-edge technologies of societal relevance and significance. This project establishes the NSF Industry/University Collaborative Research Center for Big Learning (CBL) at University of Florida (UF). With substantial breakthroughs in multiple modalities of challenges, such as computer vision, speech recognitions, and natural language understanding, the renaissance of machine intelligence is dawning. The CBL vision is to create intelligence towards intelligence-driven society. The mission is to pioneer novel deep learning algorithms, systems, and applications through unified and coordinated efforts in the CBL consortium. The UF Site will focus on intelligent platforms and applications and closely collaborate with other sites on deep learning algorithms, systems, and applications. The CBL will have broad transformative impacts in technologies, education, and society. CBL aims to create pioneering research and applications to address a broad spectrum of real-world challenges, making significant contributions and impacts to the deep learning community. The discoveries from CBL will make significant contributions to promote products and services of industry in general and CBL industry partners in particular. As the magnet of deep learning research and applications, CBL offers an ideal platform to nurture next-generation talents through world-class mentors from both academia and industry, disseminates the cutting-edge technologies, and facilitates industry/university collaboration and technology transfer. The center repository will be hosted at http://nsfcbl.org. The data, code, documents will be well organized and maintained on the CBL servers for the duration of the center for more than five years and beyond. The internal code repository will be managed by GitLab. After the software packages are well documented and tested, they will be released and managed by popular public code hosting services, such as GitHub and Bitbucket.

Human emotions are dynamic, multidimensional responses to challenges and opportunities, which emerge from network interactions in the brain. Disruptions of these network interactions underlie emotional dysregulation in many mental disorders, including anxiety and depression. Creating an AI-based model system that is informed and validated by known biological findings and can be used to carry out causal manipulations and test the consequences against human imaging data will thus be a highly significant development in the short term. The long-term goal is to understand how the human brain processes emotional information and how the process breaks down in mental disorders. NIH currently funds the team to record and analyze fMRI data from humans viewing natural images of varying emotional content. In the process of their research, they recognize that empirical studies such as theirs have significant limitations. Chief among them is the lack of ability to manipulate the system to establish the causal basis for the observed relationship between brain and behavior. The advent of AI, especially deep neural networks (DNNs), opens a new avenue to address this problem. Creating an AI-based model system that is informed and validated by known biological findings and that can be used to carry out causal manipulations and allow the testing of the consequences against human imaging data will thus be a significant step toward achieving our long-term goal.

Human emotions are dynamic, multidimensional responses to challenges and opportunities, which emerge from network interactions in the brain. Disruptions of these network interactions underlie emotional dysregulation in many mental disorders, including anxiety and depression. Creating an AI-based model system that is informed and validated by known biological findings and can be used to carry out causal manipulations and test the consequences against human imaging data will thus be a highly significant development in the short term. The long-term goal is to understand how the human brain processes emotional information and how the process breaks down in mental disorders. NIH currently funds the team to record and analyze fMRI data from humans viewing natural images of varying emotional content. In the process of their research, they recognize that empirical studies such as theirs have significant limitations. Chief among them is the lack of ability to manipulate the system to establish the causal basis for the observed relationship between brain and behavior. The advent of AI, especially deep neural networks (DNNs), opens a new avenue to address this problem. Creating an AI-based model system that is informed and validated by known biological findings and that can be used to carry out causal manipulations and allow the testing of the consequences against human imaging data will thus be a significant step toward achieving our long-term goal.

Vision-threatening diseases are one of the leading causes of blindness. DR, a common complication of diabetes, is the leading cause of blindness in American adults and the fastest growing disease threatening nearly 415 million diabetic patients worldwide. With professional eye imaging devices such as fundus cameras or Optical Coherence Tomography (OCT) scanners, most of the vision-threatening diseases can be curable if detected early. However, these diseases are still damaging people’s vision and leading to irreversible blindness, especially in rural areas and low-income communities where professional imaging devices and medical specialists are not available or not even affordable. There is an urgent need for early detection of vision-threatening diseases before vision loss in these areas. The current practice of DR screening relies on human experts to manually examine and diagnose DR in stereoscopic color fundus photographs at hospitals using professional fundus camera, which is time-consuming and infeasible for large-scale screening. It also puts an enormous burden on ophthalmologists and increases waiting lists and may undermine the standards of health care. Therein, automatic DR diagnosis systems with ophthalmologist-level performance are a critical and unmet need for DR screening. Electronic Health Records (EHR) have been increasingly implemented in US hospitals. Vast amounts of longitudinal patient data have been accumulated and are available electronically in structured tables, narrative text, and images. There is an increasing need for multimodal synergistic learning methods to link different data sources for clinical and translational studies. Recent emerging of AI technologies, especially deep learning (DL) algorithms, have greatly improved the performance of automated vision-disease diagnosis systems based on EHR data. However, the current systems are unable to detect early stage of vision-diseases. On the other hand, the clinical text provides detailed diagnosis, symptoms, and other critical observations documented by physicians, which could be a valuable resource to help lesion detection from medical images. Multimodal synergistic learning is the key to linking clinical text to medical images for lesion detection. This study proposes to leverage the narrative clinical text to improve lesion level detection from medical images via clinical Natural Language Processing (NLP). The team hypothesizes that early stage vision-threatening diseases can be detected using smartphone-based fundus camera via multimodal learning integrating clinical text and images with limited lesion-level labels via clinical NLP. The ultimate goal is to improve the early detection and prevention of vision-threatening diseases among rural and low-income areas by developing a low-cost, highly efficient system that can leverage both clinical narratives and images.




In the current era of invigorating brain research, there is emerging attention in leveraging machine learning in understanding the brain, particularly exploring the brain dynamics. With the ever-increasing amount of neuroscience data, new challenges and opportunities arise for brain dynamics analysis, such as data-driven reconstruction and computer-aided diagnosis. However, there are few attempts to bridge the semantic gaps between the raw brain imaging data and the diagnosis. We will develop robust and data-driven techniques for the purpose of modeling, estimating functional parameters from the limited data brain images, and making decision support practical based on efficient direct estimation of the brain dynamics. This is an interdisciplinary research combining medical image analysis, machine learning, neuroscience, and the domain expertise.
Instructor Evaluation: 5.00/5.00, Course Evaluation: 4.75/5.00
Course Objectives: 1. Understand multimodal data mining in the biomedical domain; 2. Understand the concept, approaches, and limitations in analyzing different modalities of biomedical data. 3. Learn to use biomedical data programming libraries and skills to analyze multimodal biomedical data.
The objectives this course are: 1) Develop a proficiency in the use of computer programming (specifically, MATLAB) to analyze biomedical measurements. 2) Develop an understanding of biomedical engineering problems that require quantitative analysis and visualization.
The objectives this course are: 1) Develop a proficiency in the use of computer programming (specifically, MATLAB) to analyze biomedical measurements. 2) Develop an understanding of biomedical engineering problems that require quantitative analysis and visualization.
Instructor Evaluation: 4.63/5.00, Course Evaluation: 4.38/5.00
Course Objectives: 1. Understand multimodal data mining in the biomedical domain; 2. Understand the concept, approaches, and limitations in analyzing different modalities of biomedical data. 3. Learn to use biomedical data programming libraries and skills to analyze multimodal biomedical data.
Instructor Evaluation: 4.92/5.00, Course Evaluation: 4.70/5.00
The objectives this course are: 1) Develop a proficiency in the use of computer programming (specifically, MATLAB) to analyze biomedical measurements. 2) Develop an understanding of biomedical engineering problems that require quantitative analysis and visualization.
Instructor Evaluation: 4.83/5.00, Course Evaluation: 4.53/5.00 (Historical highest for this course)
Course Objectives: 1. Understand multimodal data mining in the biomedical domain; 2. Understand the concept, approaches, and limitations in analyzing different modalities of biomedical data. 3. Learn to use biomedical data programming libraries and skills to analyze multimodal biomedical data.
Instructor Evaluation: 4.87/5.00
The objectives this course are: 1) Develop a proficiency in the use of computer programming (specifically, MATLAB) to analyze biomedical measurements. 2) Develop an understanding of biomedical engineering problems that require quantitative analysis and visualization.
The objectives this course are: 1) Develop a proficiency in the use of computer programming (specifically, MATLAB) to analyze biomedical measurements. 2) Develop an understanding of biomedical engineering problems that require quantitative analysis and visualization.
Data Mining is the nontrivial extraction of implicit, previously unknown, and potentially useful information from data. It has gradually matured as a discipline merging ideas from statistics, machine learning, database and etc. This is an introductory course for junior/senior computer science undergraduate students on the topic of Data Mining. Topics include data mining applications, data preparation, data reduction and various data mining techniques (such as association, clustering, classification, anomaly detection).
Machine learning is concerned with the question of how to make computers learn from experience. The ability to learn is not only central to most aspects of intelligent behavior, but machine learning techniques have become key components of many software systems. For examples, machine learning techniques are used to create spam filters, to analyze customer purchase data, to understand natural language, or to detect fraudulent credit card transactions.
Unlocking brain health through multimodal digital twins and 3D vision foundation models enables comprehensive, data-driven insights for personalized diagnostics and interventions.
Dr. Ruogu Fang prepares students in a Medical AI course for a world where AI literacy is an essential part of the medical field.
Dr. Fang with a team of students and faculty members leverage deep neural networks to predict HIV.
Dr. Fang presents a workshop presentation on the impact of AI in the medical & clinical environment.
Our SMILE Lab is featured on the Forward & Up video by the College of Engineering!
Dr. Fang is interviewed by ABC Action News in a news segment on the University of Florida's supercomputer, HiPerGator.
UF, NVIDIA partner to speed brain research using AI
University of Florida researchers joined forces with scientists at NVIDIA, UF’s partner in its artificial intelligence initiative, and the OpenACC organization to significantly accelerate brain science as part of the Georgia Tech GPU Hackathon held last month.
Artificial Intelligence Prevents Dementia?
A team at the University of Florida is using targeted transcranial direct current stimulation to save memories. “It’s a weak form of electrical stimulation applied to the scalp. And this weak electric current actually has the ability to alter how the neurons behave,” continued Woods.
UF study shows artificial intelligence’s potential to predict dementia
New research published today shows that a form of artificial intelligence combined with MRI scans of the brain has the potential to predict whether people with a specific type of early memory loss will develop Alzheimer’s disease or other form of dementia.
UF researchers fight dementia using AI technology to develop new treatment
If presented with current models of electrical brain stimulation a decade ago, Dr. Adam J. Woods would have thought of a science-fiction movie plot. Artificial intelligence now pulls the fantasy out of screens and into reality.
UF researchers using artificial intelligence to develop treatment to prevent dementia
UF researchers are developing a method they hope will prevent Alzheimer’s and Dementia. Dr. Ruogu Fang and Dr. Adam Woods are working to personalize brain stimulation treatments to make them as effective as possible.
UF researchers use AI to develop precision dosing for treatment aimed at preventing dementia
UF researchers studying the use of a noninvasive brain stimulation treatment paired with cognitive training have found the therapy holds promise as an effective, drug-free approach for warding off Alzheimer’s disease and other dementias.
Our eyes may provide early warning signs of Alzheimer’s and Parkinson’s
Forget the soul — it turns out the eyes may be the best window to the brain. Changes to the retina may foreshadow Alzheimer’s and Parkinson’s diseases, and researchers say a picture of your eye could assess your future risk of neurodegenerative disease.
Eye Blood Vessels May Diagnose Parkinson's Disease
A simple eye exam combined with powerful artificial intelligence (AI) machine learning technology could provide early detection of Parkinson's disease, according to research being presented at the annual meeting of the Radiological Society of North America.
UF researchers are looking into the eyes of patients to diagnose Parkinson's Disease
With artificial intelligence (AI), researchers have moved toward diagnosing Parkinson’s disease with, essentially, an eye exam. This relatively cheap and non-invasive method could eventually lead to earlier and more accessible diagnoses.
Blood Vessels in the Eye May Diagnose Parkinson's Disease
Using an advanced machine-learning algorithm and fundus eye images, which depict the small blood vessels and more at the back of the eye, investigators are able to classify patients with Parkinson's disease compared against a control group.
Scientists Are Looking Into The Eyes Of Patients To Diagnose Parkinson’s Disease
With artificial intelligence (AI), researchers have moved toward diagnosing Parkinson's disease with, essentially, an eye exam. This relatively cheap and non-invasive method could eventually lead to earlier and more accessible diagnoses.
Eye Exam Could Lead to Early Parkinson’s Disease Diagnosis
A simple eye exam combined with powerful artificial intelligence (AI) machine learning technology could provide early detection of Parkinson’s disease, according to research being presented at the annual meeting of the Radiological Society of North America.
Simple Eye Exam With Powerful Artificial Intelligence Could Lead to Early Parkinson’s Disease Diagnosis
A simple eye exam combined with powerful artificial intelligence (AI) machine learning technology could provide early detection of Parkinson’s disease, according to research being presented at the annual meeting of RSNA.
RSNA 20: AI-Based Eye Exam Could Aid Early Parkinson’s Disease Diagnosis
A simple eye exam combined with powerful artificial intelligence (AI) machine learning technology could provide early detection of Parkinson’s disease, according to research being presented at the annual meeting of the Radiological Society of North America.
Fang selected to ACM's Future of Computer Academy
Dr. Ruogu Fang, incoming assistant professor in the J. Crayton Pruitt Family Department of Biomedical Engineering, has been selected as a member of the Association for Computing Machinery’s (ACM) inaugural Future Computing Academy (FCA).
We are hiring a Postdoc Researcher. Proficiency with Python and deep learning is required. Experience with Pytorch/Tensorflow, FSL/FreeSurfer/SPM are preferred. Please check out the job HERE.
Please fill out this Online Form AND send me an email at smilefanglab@gmail.com with the following:
For Ph.D. applicants interested in working in my lab, you can apply for the Ph.D. program in Biomedical Engineering, Electrical and Computer Engineering, Computer Science, or other related programs at UF.
Please fill out this Online Form AND send me an email at smilefanglab@gmail.com with the following before you apply:
If you are already a master student or have received admission to BME/ECE/CISE or related programs at the University of Florida, first complete the application form below.
Please fill out this Online Form AND send me an email at smilefanglab@gmail.com with the intention for voluntary research along with the following:
Note: We are currently hiring for volunteer positions only.
We take undergraduates from all years and prefer students who have a good math and coding background, or at least the strong motivation and interest to build a solid math and coding background.
This form is always open, but the form will only be reviewed during the recruitment period. The official recruitment period begins at the end of Fall and Spring semester. More details are listed below.
Prerequisite: Complete Coursera Machine Learning course by Andrew Ng before you can officially join the lab. (Free)
Co-requisite: Students who have joined our lab (including PhD, MS, and UG) are strongly encouraged to complete the following courses online during their time in the lab:
All students that meet these requirements are welcome to apply! We are particularly interested in broadening participation of underrepresented groups in STEM, and women and minorities are especially encouraged to apply. We also aim to provide research experiences for students who have limited exposure and opportunities to participate in research.
The University of Florida (UF) is recognized as one of the nation’s leading public universities, achieving a Top 5 ranking in the recent five years by U.S. News & World Report. Located in Gainesville, a vibrant city in northern Florida, UFL is celebrated for its lush campus, innovation in research, and dynamic student life. The city is known for its rich history, cultural diversity, and excellent quality of life. UFL stands out as a leader in sustainability, cutting-edge technology, and academic excellence, offering students a supportive and inclusive environment. Below is the ranking data for UFL from various ranking institutes/organizations:
| Ranking Publisher | Category | Ranking | Year |
|---|---|---|---|
| The Wall Street Journal | Top 100 leading Universities in USA | 1 | 2025 |
| Niche | Top Public Universities in America | 5 | 2025 |
| U.S. News & World Report | Top Public Universities | 7 | 2024 |
| Forbes | Top Colleges 2025 | 26 | 2024 |
| EduRank | World University Rankings | 27 | 2024 |
| U.S. News & World Report | National Universities | 30 | 2025 |
I would be happy to talk to you if you need my assistance in your research, have intention for collaboration, or need technology translation for your company. Also, welcome to join my SMILE research group. Email is the best way to get in touch with me. Please feel free to contact me!
J287 Biomedical Science Building
University of Florida
1275 Center
Dr.
Gainesville, FL, 32611.